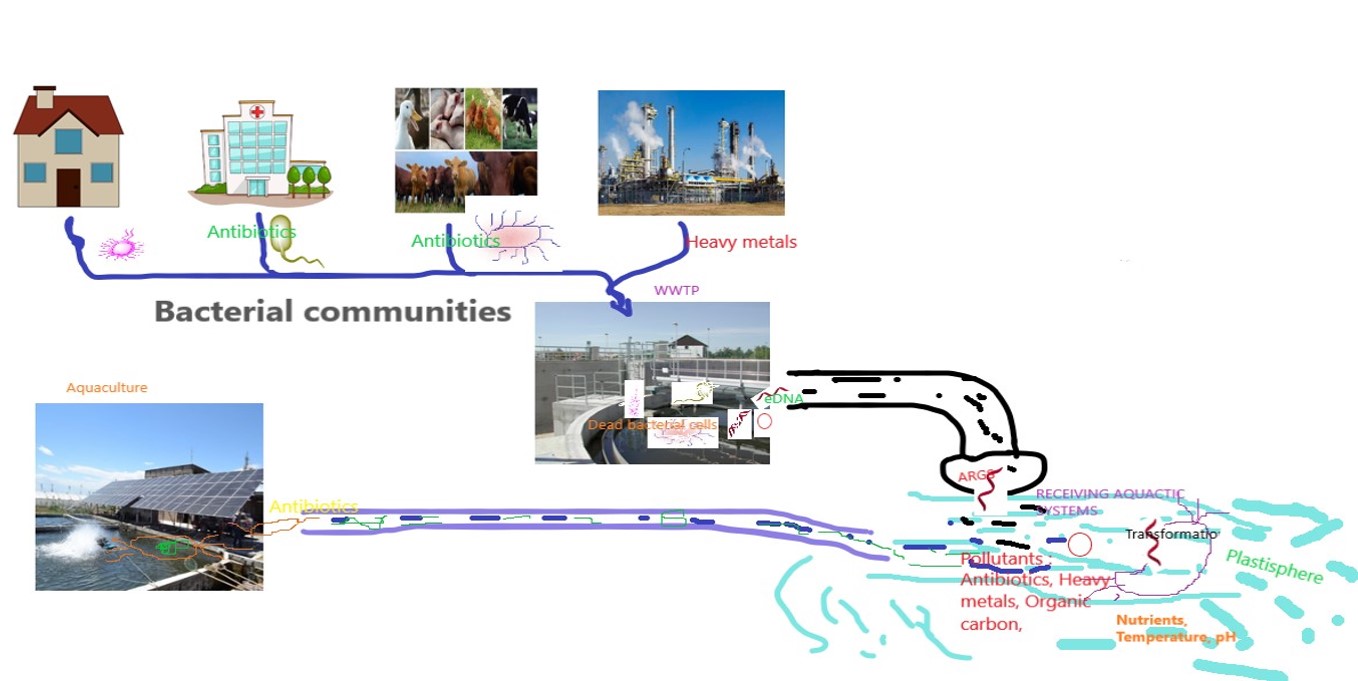
Extracellular DNA (eDNA): neglected and potential sources of antibiotic resistance genes (ARGs) in the environment
→
Europe/Rome
Description


Seminari IRSA-2022
Extracellular DNA (eDNA): neglected and potential sources of antibiotic resistance genes (ARGs) in the environment
SIVALINAM PERIYASAMY
CNR-IRSA VERBANIA
http://www.meg.irsa.cnr.it/index.php/people/post-doc-and-research-ass/sivalingam-periyasamy
In spite of antibiotic resistance being a major threat to public health, the persistence and fate of antibiotic-resistant bacteria (ARBs) and antibiotic resistance genes (ARGs) in the environment have not been comprehensively explored. Usually, ARGs occur in two forms: intracellular ARGs (iARGs) and extracellular ARGs (eARGs). Research is accumulating concerning the role of iARGs in proliferating antibiotic resistance, but there are few studies addressing the role of eARGs. Research, however, suggests that eARGs play a more crucial role in the spread of ARGs than previously thought. The focus of our work is to utilize the best available technologies to evaluate ARGs (iARGs and eARGs) and the microbial communities in aquatic environments in terms of environmental antibiotic resistome. This will facilitate the understanding of the distribution pattern of gene transfer elements and assess their abundance. Among our prime interests, we intend to test the presence of eARGs both in natural and engineered aquatic environments. The use of sequence-based metagenomic approaches will be helpful in understanding the diversity of communities and changes in an environment as a result of human-linked activities. In addition, generating sample-specific and temporal antibiotic-resistant profiles will be helpful to better understand microbial ecology and the transmission of antibiotic-resistant genes from one ecosystem to another. Also, we are interested in understanding how environmental factors such as temperature, organic matter, pH, heavy metals, and nutrients affect the persistence and spread of eARGs.
I partecipanti registrati riceveranno un link personale per connettersi mediante la piattaforma GoToMeeting.
Seminari IRSA-CNR